_78.png)
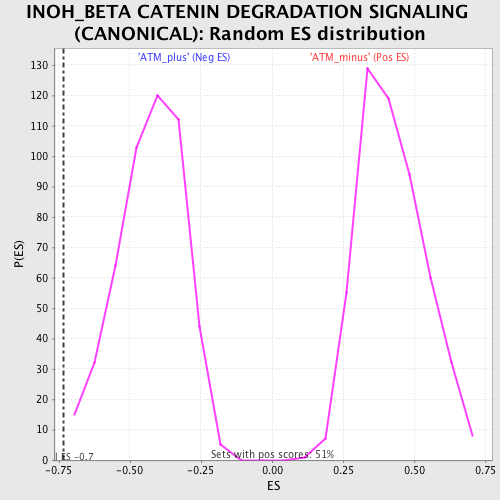
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | INOH_BETA CATENIN DEGRADATION SIGNALING (CANONICAL) |
| Enrichment Score (ES) | -0.7335278 |
| Normalized Enrichment Score (NES) | -1.7206608 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1629169 |
| FWER p-Value | 0.643 |
_78.png)
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TCF3 | 1436207_at 1450117_at | 859 | 1.448 | 0.0110 | No | ||
| 2 | YWHAB | 1420878_a_at 1420879_a_at 1420880_a_at 1436783_x_at 1438708_x_at 1455815_a_at | 1612 | 0.912 | 0.0083 | No | ||
| 3 | CTNNBIP1 | 1417567_at 1431694_a_at | 1842 | 0.815 | 0.0261 | No | ||
| 4 | DVL2 | 1417207_at 1448616_at | 2024 | 0.739 | 0.0435 | No | ||
| 5 | YWHAG | 1420816_at 1420817_at 1450012_x_at 1459693_x_at | 2251 | 0.668 | 0.0563 | No | ||
| 6 | APC | 1420956_at 1420957_at 1435543_at 1450056_at | 2651 | 0.562 | 0.0576 | No | ||
| 7 | DVL3 | 1420457_at | 4441 | 0.318 | -0.0130 | No | ||
| 8 | PYGO1 | 1431091_at | 6383 | 0.220 | -0.0940 | No | ||
| 9 | TCF7 | 1433471_at 1450461_at | 11544 | 0.069 | -0.3270 | No | ||
| 10 | NLK | 1419112_at 1435970_at 1443279_at 1445427_at 1456467_s_at | 13868 | 0.007 | -0.4328 | No | ||
| 11 | SFN | 1448612_at | 14226 | -0.004 | -0.4489 | No | ||
| 12 | FRAT2 | 1455220_at | 14829 | -0.027 | -0.4755 | No | ||
| 13 | CUL1 | 1448174_at 1459516_at | 15156 | -0.039 | -0.4890 | No | ||
| 14 | CTNNB1 | 1420811_a_at 1430533_a_at 1450008_a_at | 18001 | -0.278 | -0.6092 | No | ||
| 15 | DVL1 | 1437301_a_at 1450978_at | 19023 | -0.470 | -0.6395 | No | ||
| 16 | LEF1 | 1421299_a_at 1436398_at 1440760_at | 21085 | -1.534 | -0.6804 | Yes | ||
| 17 | TCF7L2 | 1425229_a_at 1426639_a_at 1429427_s_at 1429428_at 1441756_at 1446452_at 1447071_at | 21193 | -1.692 | -0.6266 | Yes | ||
| 18 | CREBBP | 1434633_at 1435224_at 1436983_at 1444856_at 1457641_at 1459804_at | 21262 | -1.794 | -0.5675 | Yes | ||
| 19 | AXIN1 | 1426966_at 1426967_at 1441029_at | 21350 | -1.918 | -0.5049 | Yes | ||
| 20 | AXIN2 | 1421341_at 1436845_at | 21387 | -1.982 | -0.4379 | Yes | ||
| 21 | BCL9 | 1445859_at 1446767_at 1451574_at 1455565_at | 21515 | -2.210 | -0.3670 | Yes | ||
| 22 | SMAD4 | 1422485_at 1422486_a_at 1422487_at 1444205_at | 21572 | -2.354 | -0.2880 | Yes | ||
| 23 | GSK3B | 1434439_at 1437001_at 1439931_at 1439949_at 1451020_at 1454958_at | 21811 | -3.432 | -0.1798 | Yes | ||
| 24 | YWHAE | 1426384_a_at 1426385_x_at 1435702_s_at 1438839_a_at 1440841_at 1457173_at | 21918 | -5.349 | 0.0008 | Yes |
_79.png)